DiNetxify
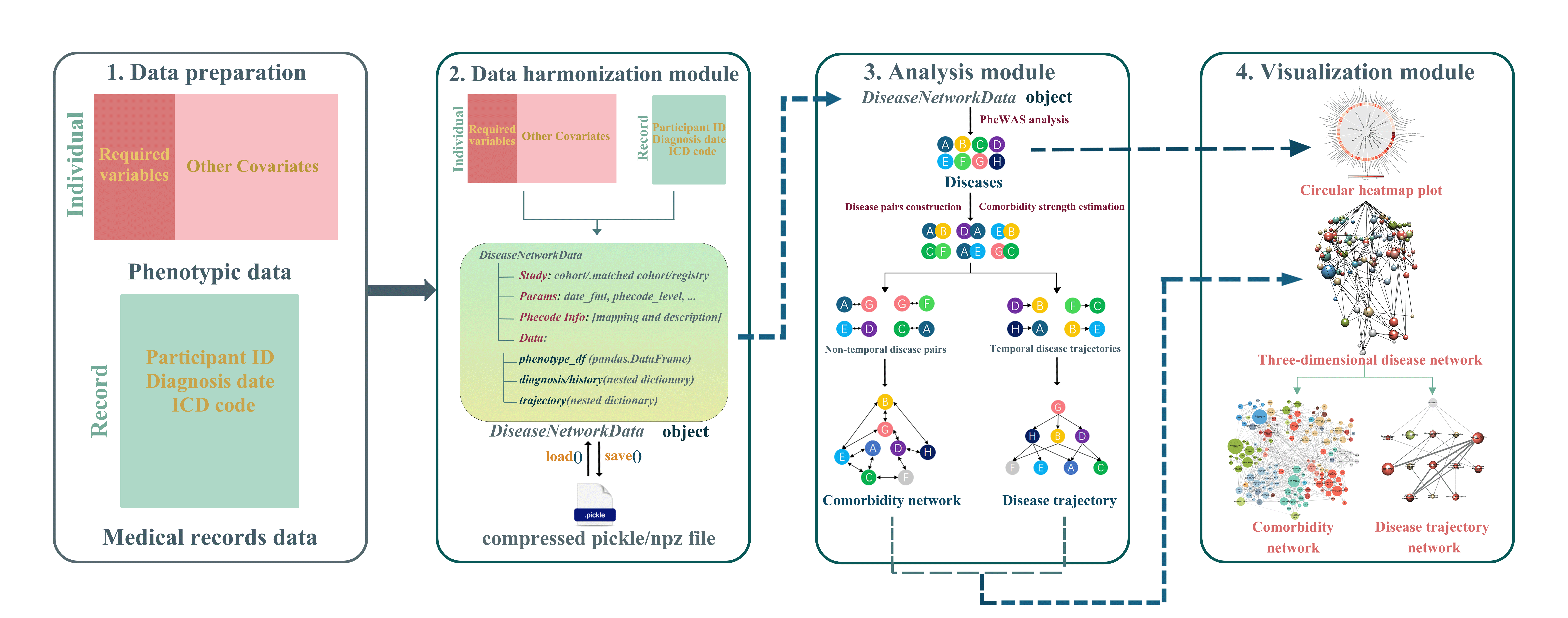
DiNetxify is an open-source Python package for comprehensive three-dimensional (3D) disease network analysis of large-scale electronic health record (EHR) data. It integrates data harmonization, statistical analysis, and visualization in a single workflow to uncover multimorbidity patterns and disease progression pathways. The package supports cohort, matched cohort, and exposed-only cohort study designs, provides both step-by-step and one-step analysis workflows, and is released under the GPL-3.0 license.
Installation
DiNetxify requires Python 3.10+. Install the latest release from PyPI with:
pip install dinetxify
Core dependencies include numpy, pandas, scipy, statsmodels, lifelines, matplotlib, plotly, networkx, python_louvain, scikit_learn, and tqdm.
Source code and issue report
Source code, release history, and issue tracking are available on GitHub: HZcohort/DiNetxify.
Citation
If you use this software in your research, please cite:
DiNetxify: a python package for three-dimensional disease network analysis based on electronic health record data (PMID: 41579291)
Disease clusters and their genetic determinants following a diagnosis of depression: analyses based on a novel three-dimensional disease network approach (PMID: 40681841)
Contact
Can Hou: houcan@wchscu.cn
Haowen Liu: haowenliu81@gmail.com